How the new HypoMap can support your research
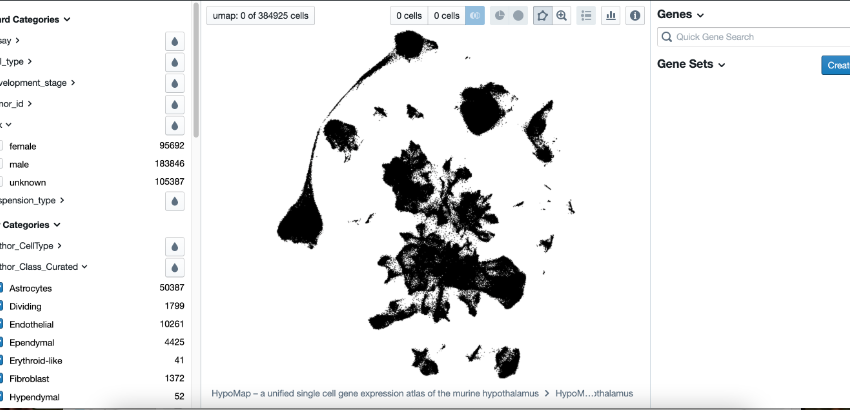
The HypoMap is a unified single cell gene expression atlas of the mouse hypothalamus and the most comprehensive database of its kind.
Launched in October 2022 in collaboration with the Max Planck Institute for Metabolism Research in Germany, Brian Lam, past BSN President Giles Yeo and colleagues from the Coll, O’Rahilly, Yeo (COY) research group at the University of Cambridge have merged 18 large mouse single-cell datasets into an integrated reference atlas of the mouse hypothalamus which they have called ‘HypoMap’.
The HypoMap is published in Nature Metabolism and is freely available to neuroendocrinologists to use in their research.
VIEW THE HYPOMAP
The British Society for Neuroendocrinology (BSN) spoke to past President Professor Giles Yeo on the newly created Hypomap.
Why did you create the HypoMap?
The hypothalamus is not just an homogenous group of cells. It is probably one of the most complex regions of the brain encompassing a number of different nuclei each containing hundreds if not thousands neuronal and other cell types. It acts broadly as a fuel sensor. The research community have been trying to understand the complexity and heterogeneity of the hypothalamus by using single cell approaches in mice. There are 10-100s of single cell data sets out there. The issue is that they are not directly comparable which is where the HypoMap comes in.
What is the HypoMap?
We have taken 18 of the largest mouse single cell datasets of the hypothalamus out there and integrated them into a single database. This is just the first version of the database. These 400,000 cells probably form the largest single cell database out there and as we collect more datasets we hope the HypoMap will be the most comprehensive single cell resource in terms of mouse hypothalamic biology there is.
How can this resource help neuroendocrinologists advance their research?
The HypoMap database is fully searchable and fully open access. You can look at your favourite gene, where it's expressed in the hypothalamus and which genes it may be co-expressed with. We hope it will be a useful resource for the neuroendocrine community.

